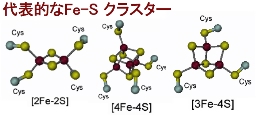
Iron-Sulfur Cluster Biosynthesis System
17/05/24 16:48
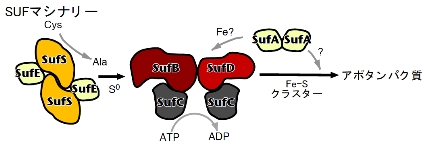
| Multicomponent enzyme machinery involved in iron-sulfur cluster biosynthesis |
| [2Fe-2S] Cluster |
[Oxidized form] 2Fe^{3+} [Reduced form] Fe^{3+} + Fe^{2+}
| [4Fe-4S] Cluster |
[Oxidized form] 2Fe^{3+} + 2Fe^{2+} [Reduced form] Fe^{3+} + 3Fe^{2+}
The
The purpose of this research is to elucidate the reaction and regulatory mechanisms of the SUF machinery: specifically, how each multi-component complex synthesizes Fe-S clusters and by what mechanism they transfer these clusters to apoproteins.
In this study, we determined the crystal structure of
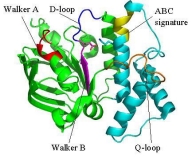
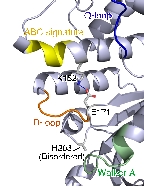
Structure and Functional Analysis of SufD with a Novel Folding Motif
The function of SufD was entirely unknown due to the lack of sequence homology with other proteins. Therefore, to obtain clues regarding its function, we determined the structure of the SufD dimer.
The SufD structure exhibits a unique fold known as a $\beta$-helix and was identified as a novel structural protein classified into a new superfamily in structural motif databases. As it was difficult to predict the function from the three-dimensional structure alone, we decided to investigate residues involved in the function using genetic methods, bearing this unique structure in mind.
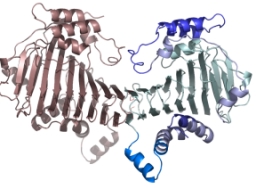
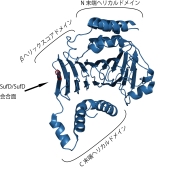
Crystal Structure of the SufD Dimer: Featuring a novel $\beta$-helix structure.
SufD Domain Structure: The dimer is formed by antiparallel $\beta$-sheets.
To investigate the structure-function relationship of SufD, we introduced mutations into amino acid residues at various sites and evaluated the in vivo function of these mutant SufDs using a complementation assay with E. coli mutant strains. This experimental system utilizes the exchange of temperature-sensitive plasmids; if SufD is non-functional, the cells cannot grow at 43°C.
The results were as follows:
- Deletion of the N-terminal $\alpha$-helical domain had almost no effect on SufD function.
- Deletion of the C-terminal $\alpha$-helical domain resulted in severe effects, and complementation ability was completely lost upon deletion of $\alpha$7–9.
- Substitution of H360 at the dimer interface with any of the other 19 amino acids resulted in loss of function.
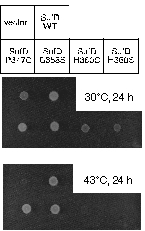

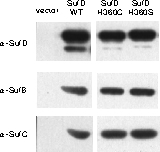
In vivo Complementation Experiment:
Introduction of a mutation at H360 in SufD renders E. coli unable to grow.
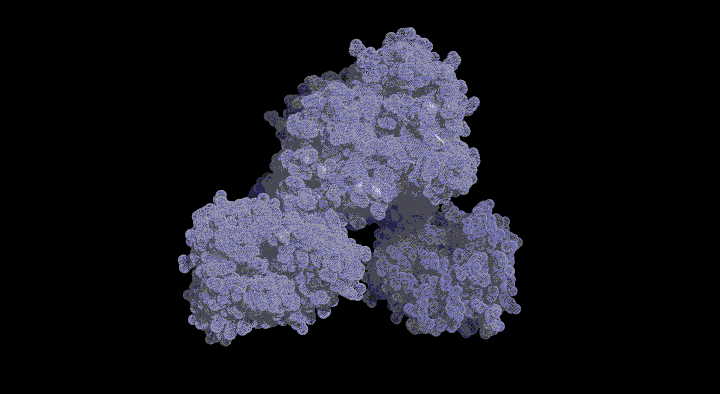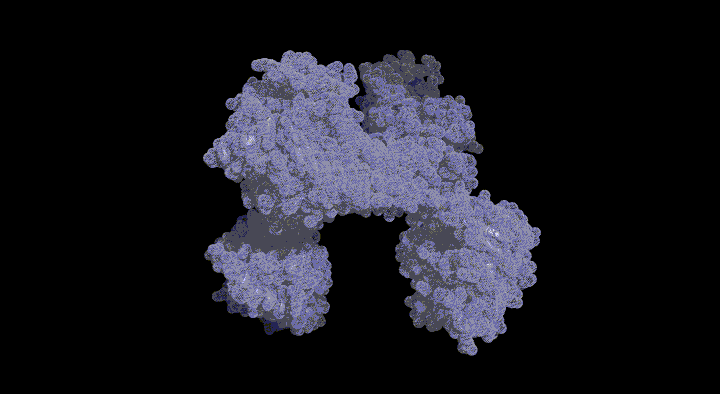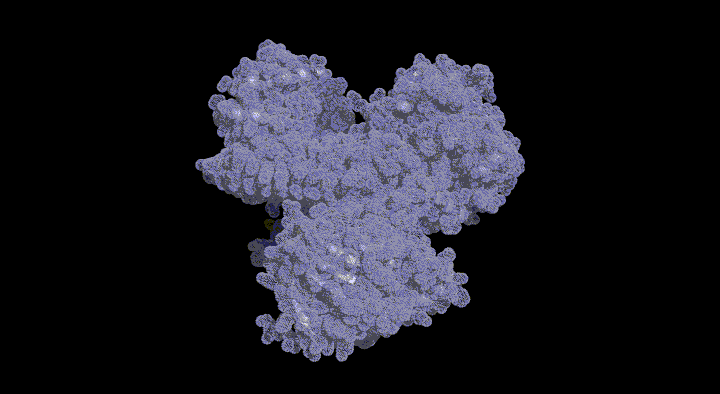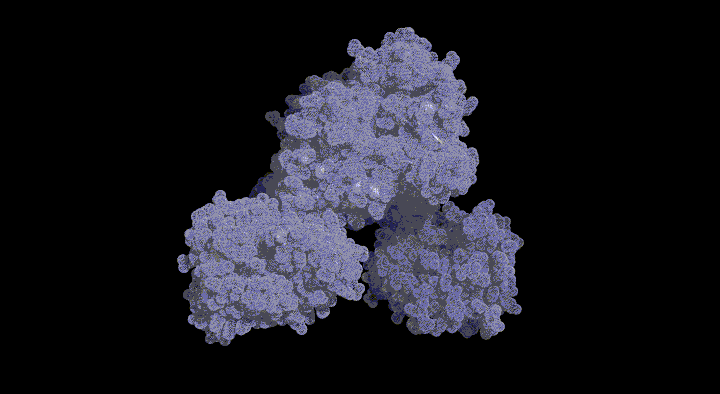
Structure of the SufC{2}-SufD{2} Complex and Structural Changes in the ATPase SufC
Upon determining the structure of the SufC${2}-SufD${2} complex, we found that SufC adopts a conformation competent for ATP hydrolysis. As mentioned previously, SufC is inactive in its monomeric state; thus, it became clear that it is activated upon complex formation.
Furthermore, the two SufC molecules bound to the SufD dimer were positioned apart from each other within the complex. Based on the analogy with ABC transporters, it is hypothesized that the two SufC molecules form a dimer.
Therefore, we introduced cysteine mutations at the predicted dimer interface of SufC to investigate whether the residues come within a distance capable of forming a disulfide bond. The results revealed that SufC within the complex forms a dimer in an ATP- and Mg$^{2+}$-dependent manner.
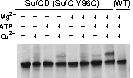
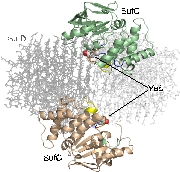
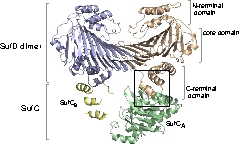
We were able to demonstrate through cross-linking experiments that the two SufC molecules within the SufC_2-SufD_2 complex form a dimer . The formation of the SufC dimer is expected to induce large conformational changes in the complex.
Operating Model of the SUF Machinery Involving Dynamic Structural Changes
Our results to date have shed light on the mechanism of Fe-S cluster biosynthesis in the SUF machinery .
We propose an operating model suggesting that the mechanism of the SUF machinery serves as the prototype for the ABC transporter superfamily—the largest in the biological world. This points to the surprising possibility that completely different functions (cluster synthesis vs. transmembrane transport) are achieved through a common underlying principle . We are currently conducting research aimed at verifying this model.
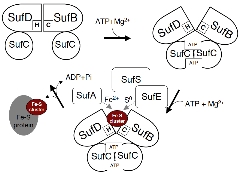
It has emerged that conformational changes in the complex induced by ATP binding expose ligands buried within the molecule (some of which have already been identified), thereby facilitating the synthesis of Fe-S clusters.