Synthesis and Degradation of Photosynthetic Pigments
17/05/24 18:40
Enzymes Involved in the Synthesis and Degradation of Photosynthetic Pigments
In photosynthetic organisms such as red algae and cyanobacteria, tetrapyrrole pigments are present in the photosynthetic apparatus and light-harvesting complexes. Efficient photosynthesis is possible only when these pigments function correctly. However, much of the metabolic pathway for the synthesis of these pigments remains to be elucidated.
I have focused my research on several key enzymes involved in pigment synthesis and degradation, aiming to uncover their functions and reaction mechanisms.
Bacteriochlorophyll c: Methylation Mechanism of the Methyltransferase Involved in Biosynthesis [J. Mol. Biol., 360, 839–849 (2006)]
Green sulfur bacteria possess chlorosomes as an extramembranous antenna system. Chlorobium tepidum contains bacteriochlorophyll c (BChl c) as the pigment within its chlorosomes. Recent genome analyses and molecular genetic approaches are beginning to reveal its biosynthetic pathway.
In this pathway, BchU was identified as the enzyme (methyltransferase) that methylates the C-20 position using S-adenosylmethionine (SAM) as a methyl donor. To date, there has been no structural information regarding methyltransferases for (bacterio)chlorophylls, and the molecular mechanism of the methylation reaction remains unknown. The aim of this study was to elucidate the structure-based SAM-dependent methylation mechanism of BchU. To this end, we performed X-ray crystal structure analysis of the enzyme.
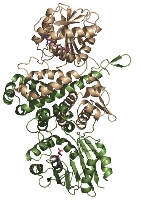
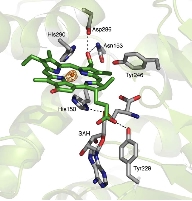
The analysis revealed that BchU forms a homodimer, with the N-terminal domain involved in dimerization. Furthermore, the C-terminal domain contained a typical Class I methyltransferase motif. We also determined the structure of the complex with SAM, clarifying its binding mode with BchU.
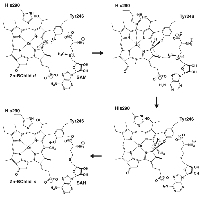
Based on these structural analyses and mutagenesis experiments, we elucidated the SAM-dependent methylation mechanism in (bacterio)chlorophylls. We are currently conducting structure-based mutagenesis experiments to further clarify the detailed substrate recognition mechanism.
Elucidation of the Reaction Mechanism of CbiL Responsible for Formation of the Basic Tetrapyrrole Skeleton of Vitamin B12
[FEBS Journal, 274, 563–573 (2007)]
Vitamin B12 is a small molecule with a complex structure, and its biosynthesis involves more than 15 enzymes. The basic skeleton of Vitamin B12 consists of a corrin ring with a central cobalt atom. This features a "distorted" structure in which one carbon atom (at the C-20 position) has been eliminated from the typical tetrapyrrole ring. This carbon elimination reaction is initiated by the methylation of the C-20 carbon by an enzyme called CbiL. The aim of this study is to elucidate the mechanism of C-20 methylation within the Vitamin B12 biosynthetic pathway.
We determined the crystal structure of CbiL in complex with a methyl donor analog (S-adenosylhomocysteine: SAH) and identified putative catalytic residues that were previously unknown. Furthermore, computer simulations predicting the binding mode of the substrate tetrapyrrole compound suggested that substrate binding is accompanied by large inter-domain conformational changes. We are currently working to experimentally verify these structure-based predictions.
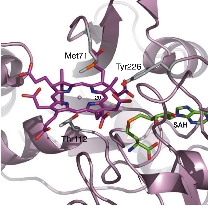
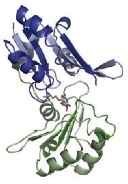
The CbiL monomer is composed of two domains, and it was revealed that the substrate binds within the cleft between them. Calculations of the domain movement (TLS motion) predicted that the two domains enclose the substrate in a "crab claw"-like manner.
-------------------------- Under Construction Below ------------------------
In photosynthetic organisms such as red algae and cyanobacteria, tetrapyrrole pigments are present in the photosynthetic apparatus and light-harvesting complexes. Efficient photosynthesis is possible only when these pigments function correctly. However, much of the metabolic pathway for the synthesis of these pigments remains to be elucidated.
I have focused my research on several key enzymes involved in pigment synthesis and degradation, aiming to uncover their functions and reaction mechanisms.
Bacteriochlorophyll c: Methylation Mechanism of the Methyltransferase Involved in Biosynthesis [J. Mol. Biol., 360, 839–849 (2006)]
Green sulfur bacteria possess chlorosomes as an extramembranous antenna system. Chlorobium tepidum contains bacteriochlorophyll c (BChl c) as the pigment within its chlorosomes. Recent genome analyses and molecular genetic approaches are beginning to reveal its biosynthetic pathway.
In this pathway, BchU was identified as the enzyme (methyltransferase) that methylates the C-20 position using S-adenosylmethionine (SAM) as a methyl donor. To date, there has been no structural information regarding methyltransferases for (bacterio)chlorophylls, and the molecular mechanism of the methylation reaction remains unknown. The aim of this study was to elucidate the structure-based SAM-dependent methylation mechanism of BchU. To this end, we performed X-ray crystal structure analysis of the enzyme.
The analysis revealed that BchU forms a homodimer, with the N-terminal domain involved in dimerization. Furthermore, the C-terminal domain contained a typical Class I methyltransferase motif. We also determined the structure of the complex with SAM, clarifying its binding mode with BchU.
Based on these structural analyses and mutagenesis experiments, we elucidated the SAM-dependent methylation mechanism in (bacterio)chlorophylls. We are currently conducting structure-based mutagenesis experiments to further clarify the detailed substrate recognition mechanism.



Elucidation of the Reaction Mechanism of CbiL Responsible for Formation of the Basic Tetrapyrrole Skeleton of Vitamin B12
[FEBS Journal, 274, 563–573 (2007)]
Vitamin B12 is a small molecule with a complex structure, and its biosynthesis involves more than 15 enzymes. The basic skeleton of Vitamin B12 consists of a corrin ring with a central cobalt atom. This features a "distorted" structure in which one carbon atom (at the C-20 position) has been eliminated from the typical tetrapyrrole ring. This carbon elimination reaction is initiated by the methylation of the C-20 carbon by an enzyme called CbiL. The aim of this study is to elucidate the mechanism of C-20 methylation within the Vitamin B12 biosynthetic pathway.
We determined the crystal structure of CbiL in complex with a methyl donor analog (S-adenosylhomocysteine: SAH) and identified putative catalytic residues that were previously unknown. Furthermore, computer simulations predicting the binding mode of the substrate tetrapyrrole compound suggested that substrate binding is accompanied by large inter-domain conformational changes. We are currently working to experimentally verify these structure-based predictions.


The CbiL monomer is composed of two domains, and it was revealed that the substrate binds within the cleft between them. Calculations of the domain movement (TLS motion) predicted that the two domains enclose the substrate in a "crab claw"-like manner.
-------------------------- Under Construction Below ------------------------